What is Spina Bifida?
|
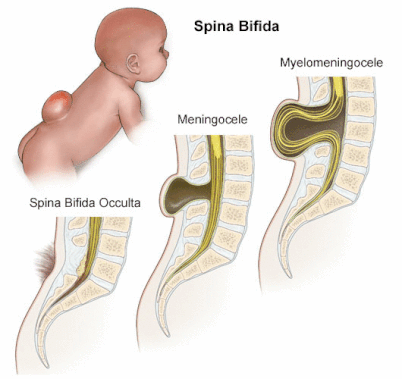
Spina Bifida is the most common birth defect in the United States. It is characterized by the failure of the ventral neural tube to fuse during development. This gap can range in severity as depicted in Figure 1. Myelomeningocele, the most severe form of spina bifida is associated with a range of symptoms including muscle weakness in the legs and lower body, possible paralysis in the lower body, bowel and bladder dysfunction, and co-morbid spinal defects, including scoliosis. Many of these defects result from nerve damage that occurs when the spinal cord bulges through the spine. |
Spina Bifida and MTHFR
Spina bifida is a multifactorial disorder. This means that many genetics and environmental factors contribute to its development. The most important environmental contribution is folate deficiency which can generally be rescued by dietary supplements. It was the discovery of this link between folate deficiency and spina bifida that uncovered the important contribution of the MTHFR gene to the development of spina bifida.
MTHFR encodes an enzyme that functions as a biological reducer in the folate metabolic pathway. Biologically, it contributes to the re-methylation of homocysteine to produce the amino acid, methionine. More specifically it reduces 5,10 methylene-tetrahydrofolate to 5 methyl-tetrahydrofolate. The enzyme acts in the plasma membrane of the cell.
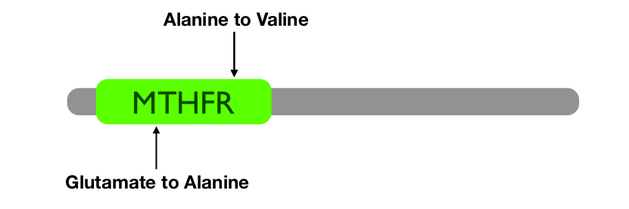
Two pathogenic mutations have been identified in humans: C677T which creates an alanine to valine substitution and A1298C which creates a glutamate to alanine change. Both of these mutations occur in the MTHFR domain.
MTHFR encodes an enzyme that functions as a biological reducer in the folate metabolic pathway. Biologically, it contributes to the re-methylation of homocysteine to produce the amino acid, methionine. More specifically it reduces 5,10 methylene-tetrahydrofolate to 5 methyl-tetrahydrofolate. The enzyme acts in the plasma membrane of the cell.
Two pathogenic mutations have been identified in humans: C677T which creates an alanine to valine substitution and A1298C which creates a glutamate to alanine change. Both of these mutations occur in the MTHFR domain.
The MTHFR Enzyme
|
Protein Interactions:
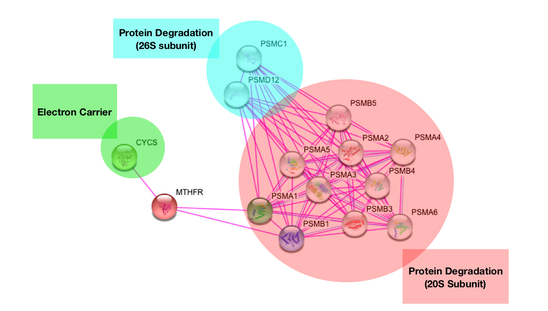
The MTHFR enzyme is known to interact with two main types of proteins. The first protein it interacts with is CYCS. This is an electron carrier protein. This interaction is logical because MTHFR has to transfer electrons to other molecules to act as a reducer. The other interaction is with the 20S subunit of the proteasome and by extent the 26S subunit of the proteasome. The proteasome functions in ubiquitin-mediated protein degradation. This makes sense because the proteasome is also implicated in cell differentiation which is important for proper development of biological structures, such as the spine. |
|
Relation of MTHFR Orthologs:
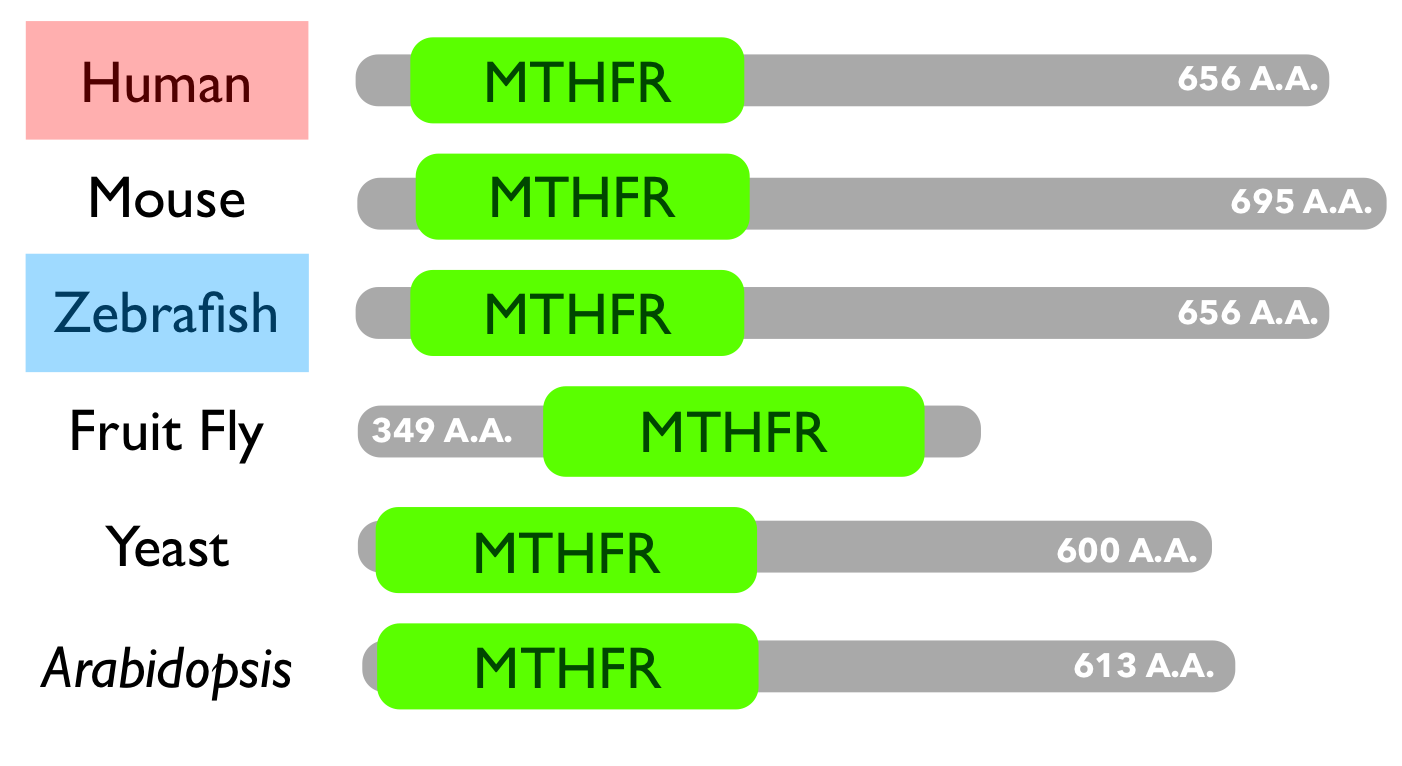
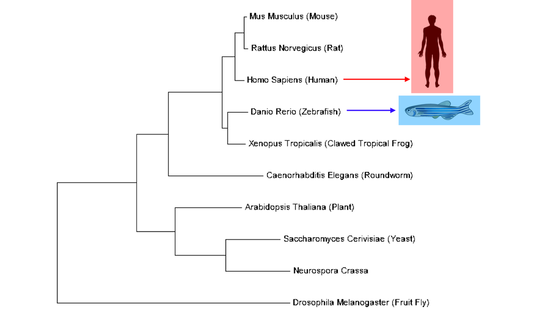
MTHFR is a well-conserved protein with orthologs present in nearly all model organisms including single-celled organisms and plants. For the purposes of this project, it is important to consider human MTHFR which is most closely related to other mammal versions of MTHFR but is also closely related to other complex vertebrates such as fish and frogs. Zebrafish will be the model organism for this project. Zebrafish MTHFR is very closely related to human MTHFR. |
|
Domain Conservation of MTHFR
MTHFR has one characterized domain, also called MTHFR, which functions as a reducer. Overall, this domain is conserved in every ortholog. Human and zebrafish MTHFR are nearly identical which suggests similar function and structure. The outlying ortholog is drosophila, or fruit fly. Its MTHFR protein is truncated, but the MTHFR domain itself is still conserved. |
Gap in Knowledge
|
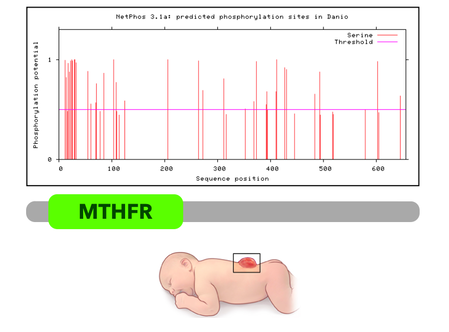
The spectrum shown to the left is a map of predicted phosphorylation sites along the MTHFR protein, created using NetPhos. It is evident that several serines at the N-terminal end of the protein are phosphorylated. However, it is not understood how regulation of MTHFR by phosphorylation affects healthy neural tube closure. |
Goals and Hypotheses
The primary goal of this project is to better understand how the regulation of MTHFR through phosphorylation affects neural tube closure. My hypothesis is that proper phosphorylation of MTHFR is important for neural tube closure and abnormal phosphorylation will lead to spina bifida. My long term goal is to understand how phosphorylation affects the larger protein interaction network of MTHFR, specifically its interactions with the proteasome which is implicated in both development and neurodegeneration.
Model Organism
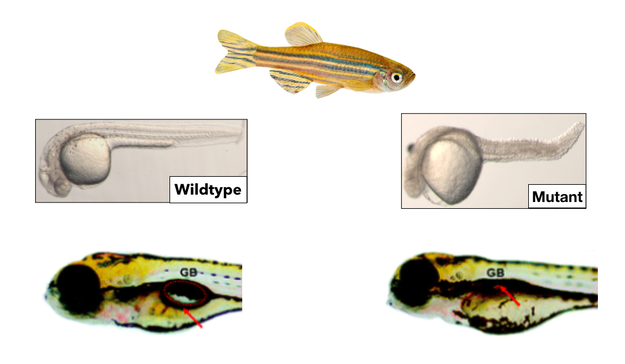
Zebrafish (Danio rerio) will be used as the model organism for this project. Zebrafish are a great model organism for practical reasons such as being relatively inexpensive and fast-developing. For example, the zebrafish neural tube closes in just 24 hours which makes it easy to study neurulation and run a lot of experiments in a short period of time. Zebrafish also display an obvious disease phenotype. When they are folate-deficient, zebrafish have a curved spine and kinked tail as well as a deflated gas bladder. These features correspond closely to the human phenotypes of muscle weakness, spine abnormalities, and bladder dysfunction, making zebrafish an excellent model for spina bifida.
Aim 1
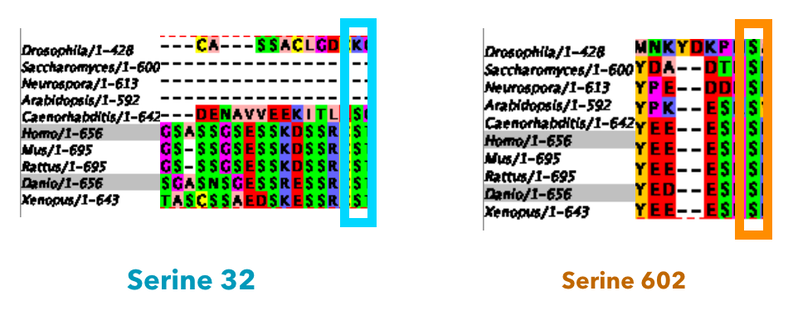
My first goal will be to identify serines important to this project. This will be accomplished by using Blast to identify protein sequences then using a website called Clustal Omega to align these sequences. I will be looking first for a serine that is conserved in vertebrates only and is located in the heavily phosphorylated N-terminal region of the MTHFR domain. For the purposes of this project, I will use serine 32. I will then look for a serine conserved in all species with MTHFR homologs which will be serine 602 for the purposes of this project. This will serve as a control mutant. This is important because MTHFR is involved with a lot of processes, and I am only interested in effects specific to the development of a spine.
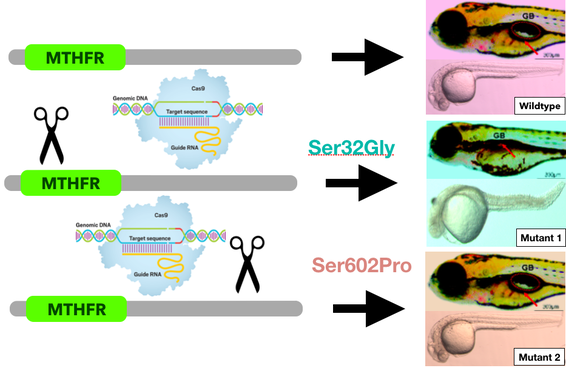
I will then mutate these serine sites using CRISPR CAS9. I will change the serine 32 to a glycine residue and the serine 602 (control mutant) to a proline residue. This will create three populations of fish, the serine 32 mutant, the serine 602 control mutant, and wild type fish.
I hypothesize that the serine 32 mutants will exhibit the disease phenotype with a curved spine and deflated swim bladder. The control mutants will not have these features although they may have other off-target phenotypic defects. Wild type fish are expected to develop completely normally.
Aim 2
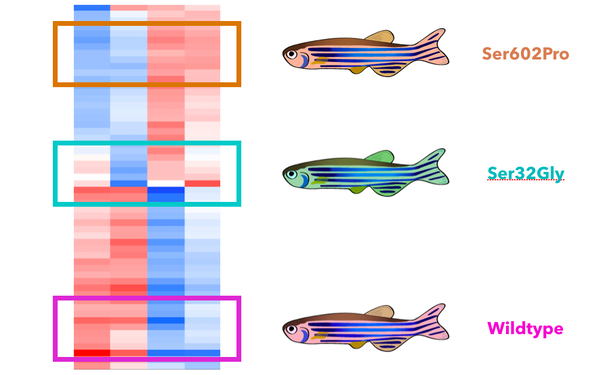
My next goal is to understand the role of MTHFR phosphorylation in regulating gene expression. I will run a micrroarray to analyze which genes are differentially regulated as a result of aberrant MTHFR phosphorylation. It is expected that all three fish populations will show different heat map patterns. It will be important to isolate specifically the genes that are differentially expressed from the wild type in the serine 32 mutants but not in the serine 602 mutants. These genes are most likely specific to neurulation. Gene ontology will then be used to characterize the processes that these specific genes affect.
I hypothesize that the genes differentially regulated in the serine 32 mutants exclusively will most likely be involved with developmental processes like cell differentiation, division, and migration. I expect the genes differentially expressed in both mutants will likely be more general to metabolism.
Aim 3
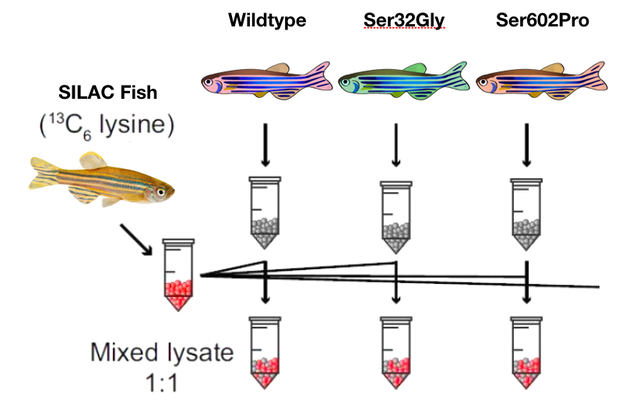
My final goal will be to quantify phosphorylation in both mutant fish versus wildtype fish. This will be accomplished using SILAC and mass spectrometry. I will quantify phosphorylation sites up and down regulated in both mutants as compared to the wild type fish. I will again be most interested in the differentially regulated sites exclusively identified in the serine 32 mutants. Gene ontology will then be used to categorize the processes affected.
I hypothesize that there will be significant increases in phosphorylation of proteins involved with spinal and structural development in the serine32 mutants but not the control mutants.
Future Directions
In the future, it will be important to try to understand the role of phosphorylation of MTHFR in the context of its protein interaction network, specifically the proteasome. The proteasome is important for processes like cell differentiation during development, but it is also implicated in neurodegenerative diseases like Alzheimer's and Parkinson's when is it not functioning normally. Understanding the role of phosphorylation and protein regulation within this network will be imperative to work toward understanding the human nervous system and its function normally and abnormally.
Final Talk and Drafts
Rogers_FinalDraft_4.23_PDF
Rogers_FinalDraft_4.23_Keynote
Rogers_DraftOne_PowerPoint
Rogers_DraftOne_KeyNote
Rogers_FinalDraft_4.23_Keynote
Rogers_DraftOne_PowerPoint
Rogers_DraftOne_KeyNote
References (Final Talk and Conclusions Page):
1. Araya, C., Ward, L., Girdler, G.C. & Miranda, M. (2016). Coordinating cell and tissue behavior during zebrafish neural tube morphogenesis. Developmental Dynamics, 245, 197-208.
2. Ben-Nissan, G., Sharon, M. (2014). Regulating the 20S proteome ubiquitin-independent degradation pathway. Biomolecules, 4, 862-884.
3. Beaudin, A. & Stover, P.J. (2009). Insights into metabolic mechanisms underlying folate-responsive neural tube defects: A minireview. Clinical and Molecular Teratology, 85, 274-284.
4. Copp, A.J., Adzick, N.S., Chitty, L.S., Fletcher, J.M., Holmbeck, G.N. & Shaw, G.M. (2016). Spina bifida. Nature Reviews Disease Primers.
5. Demassi, M., Netto, L.E.S., Silva, G.M., Hand, A., Oliviera, C.L.P, Bicev, G.N., Gozzo, F., Barros, M.H., Leme, J.M.M. & Ohara, E. (2014). Redox regulation of the proteosome via S-glutathionylation. Redox Biology. 2, 44-51.
6. Greco, M., Chiriaco, F., Del Boccio, P., Tagliaferro, L., Acierno, R., MEnegazzi, P., Pinca, E., Pignatelli, F., Storelli, C., Federici, G., Urbani, A. & Maffia, M. (2006). A proteomic approach for the characterization of C677T mutation of human gene methylenetetrahydrofolate reductase. Proteomics, 6, 5350-5360.
7. Lee, M.S., Bonner, J.R., Bernard, D.J., Sanchez, R.L., Sause, E.T., Prentice, R.R., Burgess, S.M. & Brody, L.C. (2012). Disruption of the folate pathway in zebrafish causes developmental defects. BMC Developmental Biology, 12.
8. Marini, N.J., Gin, J., Ziegle, J., Keho, K.., Ginzinger, D., Gilbert, D.A., Rine, J. (2008). The prevalence of folate-remedial MTHFR enzyme variants in humans. Biological Sciences, 105, 8055-8060.
9. National Institutes of Health. (n.d.). MTHFR Gene. Retrieved from https://ghr.nlm.nih.gov/gene/MTHFR#location
10. NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2013;41:D8–D20. [PMC free article] [PubMed]
11. Obeid, R. & Herrmann, W. (2006). Mechanisms of homocysteine neurotoxicity in neurodegenerative diseases with special reference to dementia. Federation of European Biochemical Sciences Press, 580.
12. Van Ham, T.J., Mapes, J., Kokel, D., Peterson, R.T. (2019). Live imaging of apoptotic cells in zebrafish. The FASEB Journal, 24, 4336-4342.
13. Zhang, T., Lou, J., Zhong, R., Wu, J., Zou, L., Sun, Y., Lu, X., Liu, L., Miao, X. & Xiong, G. (2013). Genetic variants in the folate pathway and the risk of neural tube defects: A meta-analysis of the published literature. PLOS One, 8, e59570.
Images:
Header: https://www.labroots.com/trending/microbiology/5278/superspreader-phages-transfer-drug-resistance
https://www.medicalnewstoday.com/articles/220424.php
https://issuu.com/choppublications/docs/spina-bifida-flip-book/3
http://flipper.diff.org/app/items/info/6483
https://www.stanfordchildrens.org/en/topic/default?id=spina-bifida-90-P02415
https://www.everydayhealth.com/g00/multiple-sclerosis/symptoms/coping-with-bladder-dysfunction/?i10c.ua=1&i10c.encReferrer=aHR0cHM6Ly93d3cuZ29vZ2xlLmNvbS8%3d&i10c.dv=21
http://www.dailymirror.lk/article/Muscle-weakness-and-Sunshine-128662.html
http://correctscoliosis.com/
https://www.health.harvard.edu/staying-healthy/the-a-list-of-b12-foods
https://www.womansday.com/health-fitness/wellness/a51060/how-to-never-be-jealous-of-20-year-olds-bodies/
https://www.rdmag.com/article/2019/03/bacteria-could-be-alternative-chemical-pesticides-crop-protection
https://curemedical.com/spina-bifida-allergies-know-products-foods/spina-bifida-kids/
https://ghr.nlm.nih.gov/gene/MTHFR#location
http://www.lazytweet.com/2018/09/person-thinking-transparent-vector/
https://www.ebi.ac.uk/interpro/potm/2006_8/Page2.htm
http://csb.cse.yzu.edu.tw/GSHSite/
https://www.bio-rad-antibodies.com/western-blotting-immunoblotting-introduction.html
https://nypost.com/2018/10/20/employees-and-execs-are-failing-drug-tests-at-shocking-rates/https://nypost.com/2018/10/20/employees-and-execs-are-failing-drug-tests-at-shocking-rates/
https://www.emdgroup.com/en/stories/use_of_fish_embryos_leads_to_less_animal_testing.html
https://string-db.org/cgi/network.pl?taskId=Q4v3DfjuYzQ0
https://www.ebi.ac.uk/Tools/services/web/toolresult.ebi?jobId=clustalo-I20190404-213834-0296-7002724-p1m&analysis=summary
https://jmg.bmj.com/content/41/6/454
https://commons.wikimedia.org/wiki/File:MTHFR_reaction.svg
https://www.proteinatlas.org/ENSG00000177000-MTHFR/cell
https://www.noldus.com/blog/how-to-mark-zebrafish-without-compromising-their-behavior
https://blast.ncbi.nlm.nih.gov/Blast.cgi
http://www.cbs.dtu.dk/services/NetPhos/
https://en.wikipedia.org/wiki/File:Scissors_icon_black.svg
http://www.njsta.org/news/crispr-in-the-classroom-by-simon-levien
https://www.researchgate.net/figure/The-heat-map-depicts-unsupervised-clustering-of-data-from-expression-arrays-from-two_fig2_235940973
https://www.pinnaclehealth.org/wellness-library/blog-and-healthwise/wellness-collection/article/en-us/tp12464/
1. Araya, C., Ward, L., Girdler, G.C. & Miranda, M. (2016). Coordinating cell and tissue behavior during zebrafish neural tube morphogenesis. Developmental Dynamics, 245, 197-208.
2. Ben-Nissan, G., Sharon, M. (2014). Regulating the 20S proteome ubiquitin-independent degradation pathway. Biomolecules, 4, 862-884.
3. Beaudin, A. & Stover, P.J. (2009). Insights into metabolic mechanisms underlying folate-responsive neural tube defects: A minireview. Clinical and Molecular Teratology, 85, 274-284.
4. Copp, A.J., Adzick, N.S., Chitty, L.S., Fletcher, J.M., Holmbeck, G.N. & Shaw, G.M. (2016). Spina bifida. Nature Reviews Disease Primers.
5. Demassi, M., Netto, L.E.S., Silva, G.M., Hand, A., Oliviera, C.L.P, Bicev, G.N., Gozzo, F., Barros, M.H., Leme, J.M.M. & Ohara, E. (2014). Redox regulation of the proteosome via S-glutathionylation. Redox Biology. 2, 44-51.
6. Greco, M., Chiriaco, F., Del Boccio, P., Tagliaferro, L., Acierno, R., MEnegazzi, P., Pinca, E., Pignatelli, F., Storelli, C., Federici, G., Urbani, A. & Maffia, M. (2006). A proteomic approach for the characterization of C677T mutation of human gene methylenetetrahydrofolate reductase. Proteomics, 6, 5350-5360.
7. Lee, M.S., Bonner, J.R., Bernard, D.J., Sanchez, R.L., Sause, E.T., Prentice, R.R., Burgess, S.M. & Brody, L.C. (2012). Disruption of the folate pathway in zebrafish causes developmental defects. BMC Developmental Biology, 12.
8. Marini, N.J., Gin, J., Ziegle, J., Keho, K.., Ginzinger, D., Gilbert, D.A., Rine, J. (2008). The prevalence of folate-remedial MTHFR enzyme variants in humans. Biological Sciences, 105, 8055-8060.
9. National Institutes of Health. (n.d.). MTHFR Gene. Retrieved from https://ghr.nlm.nih.gov/gene/MTHFR#location
10. NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2013;41:D8–D20. [PMC free article] [PubMed]
11. Obeid, R. & Herrmann, W. (2006). Mechanisms of homocysteine neurotoxicity in neurodegenerative diseases with special reference to dementia. Federation of European Biochemical Sciences Press, 580.
12. Van Ham, T.J., Mapes, J., Kokel, D., Peterson, R.T. (2019). Live imaging of apoptotic cells in zebrafish. The FASEB Journal, 24, 4336-4342.
13. Zhang, T., Lou, J., Zhong, R., Wu, J., Zou, L., Sun, Y., Lu, X., Liu, L., Miao, X. & Xiong, G. (2013). Genetic variants in the folate pathway and the risk of neural tube defects: A meta-analysis of the published literature. PLOS One, 8, e59570.
Images:
Header: https://www.labroots.com/trending/microbiology/5278/superspreader-phages-transfer-drug-resistance
https://www.medicalnewstoday.com/articles/220424.php
https://issuu.com/choppublications/docs/spina-bifida-flip-book/3
http://flipper.diff.org/app/items/info/6483
https://www.stanfordchildrens.org/en/topic/default?id=spina-bifida-90-P02415
https://www.everydayhealth.com/g00/multiple-sclerosis/symptoms/coping-with-bladder-dysfunction/?i10c.ua=1&i10c.encReferrer=aHR0cHM6Ly93d3cuZ29vZ2xlLmNvbS8%3d&i10c.dv=21
http://www.dailymirror.lk/article/Muscle-weakness-and-Sunshine-128662.html
http://correctscoliosis.com/
https://www.health.harvard.edu/staying-healthy/the-a-list-of-b12-foods
https://www.womansday.com/health-fitness/wellness/a51060/how-to-never-be-jealous-of-20-year-olds-bodies/
https://www.rdmag.com/article/2019/03/bacteria-could-be-alternative-chemical-pesticides-crop-protection
https://curemedical.com/spina-bifida-allergies-know-products-foods/spina-bifida-kids/
https://ghr.nlm.nih.gov/gene/MTHFR#location
http://www.lazytweet.com/2018/09/person-thinking-transparent-vector/
https://www.ebi.ac.uk/interpro/potm/2006_8/Page2.htm
http://csb.cse.yzu.edu.tw/GSHSite/
https://www.bio-rad-antibodies.com/western-blotting-immunoblotting-introduction.html
https://nypost.com/2018/10/20/employees-and-execs-are-failing-drug-tests-at-shocking-rates/https://nypost.com/2018/10/20/employees-and-execs-are-failing-drug-tests-at-shocking-rates/
https://www.emdgroup.com/en/stories/use_of_fish_embryos_leads_to_less_animal_testing.html
https://string-db.org/cgi/network.pl?taskId=Q4v3DfjuYzQ0
https://www.ebi.ac.uk/Tools/services/web/toolresult.ebi?jobId=clustalo-I20190404-213834-0296-7002724-p1m&analysis=summary
https://jmg.bmj.com/content/41/6/454
https://commons.wikimedia.org/wiki/File:MTHFR_reaction.svg
https://www.proteinatlas.org/ENSG00000177000-MTHFR/cell
https://www.noldus.com/blog/how-to-mark-zebrafish-without-compromising-their-behavior
https://blast.ncbi.nlm.nih.gov/Blast.cgi
http://www.cbs.dtu.dk/services/NetPhos/
https://en.wikipedia.org/wiki/File:Scissors_icon_black.svg
http://www.njsta.org/news/crispr-in-the-classroom-by-simon-levien
https://www.researchgate.net/figure/The-heat-map-depicts-unsupervised-clustering-of-data-from-expression-arrays-from-two_fig2_235940973
https://www.pinnaclehealth.org/wellness-library/blog-and-healthwise/wellness-collection/article/en-us/tp12464/