This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What Are Protein-Protein Interactions?
Protein-protein interactions are specific physical or chemical interactions between proteins. These interactions lead to the formation of complexes which perform specific functions. Most biological functions occur as a result of protein interactions rather than proteins acting alone. This makes these interactions incredibly important to understand. The interactome is the term that refers to all of the protein interactions occurring within an organism. [1]
The MTHFR Interaction Network
Databases such as String, IntAct, and BioGRID can be used to search for protein interactions that have been empirically demonstrated. The MTHFR interaction network was investigated using String.
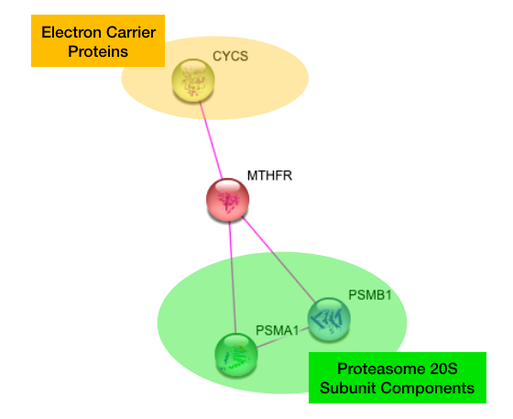
Human MTHFR
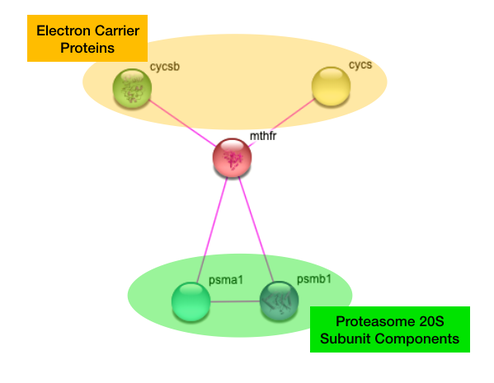
Zebrafish MTHFR
Discussion
The human and zebrafish MTHFR interaction networks are identical. It is not surprising that MTHFR interacts with Cytochrome C in humans and its equivalent two copies in zebrafish because cytochrome c is an electron-carrying protein, and MTHFR functions as a reducer.
However, the interaction of MTHFR with components of the 20S subunit of the central proteasome was quite surprising. The central proteasome engages in ubiquitin-mediated protein degradation and is necessary for proper cell differentiation. [2]
The function of the central proteasome requires a reducing environment. [3] I hypothesize that reduction in the reducing function provided by MTHFR may decrease proteasome functioning. This would in turn decrease cell differentiation and interfere with the fusing of the neural tube.
However, the interaction of MTHFR with components of the 20S subunit of the central proteasome was quite surprising. The central proteasome engages in ubiquitin-mediated protein degradation and is necessary for proper cell differentiation. [2]
The function of the central proteasome requires a reducing environment. [3] I hypothesize that reduction in the reducing function provided by MTHFR may decrease proteasome functioning. This would in turn decrease cell differentiation and interfere with the fusing of the neural tube.
References:
[1] De Las Rivas, J. & Fontanillo, C. (2010). Protein-protein interactions essentials: Key concepts to building and analyzing interactome networks. PLOS Computational Biology, 6, e10000807.
[2] Ben-Nissan, G., Sharon, M. (2014). Regulating the 20S proteome ubiquitin-independent degradation pathway. Biomolecules, 4, 862-884.
[3] Demassi, M., Netto, L.E.S., Silva, G.M., Hand, A., Oliviera, C.L.P, Bicev, G.N., Gozzo, F., Barros, M.H., Leme, J.M.M. & Ohara, E. (2014). Redox regulation of the proteosome via S-glutathionylation. Redox Biology. 2, 44-51.
Images:
Header: https://www.vocus.co.nz/our-network
Images 2 and 3: https://string-db.org/cgi/input.pl
[1] De Las Rivas, J. & Fontanillo, C. (2010). Protein-protein interactions essentials: Key concepts to building and analyzing interactome networks. PLOS Computational Biology, 6, e10000807.
[2] Ben-Nissan, G., Sharon, M. (2014). Regulating the 20S proteome ubiquitin-independent degradation pathway. Biomolecules, 4, 862-884.
[3] Demassi, M., Netto, L.E.S., Silva, G.M., Hand, A., Oliviera, C.L.P, Bicev, G.N., Gozzo, F., Barros, M.H., Leme, J.M.M. & Ohara, E. (2014). Redox regulation of the proteosome via S-glutathionylation. Redox Biology. 2, 44-51.
Images:
Header: https://www.vocus.co.nz/our-network
Images 2 and 3: https://string-db.org/cgi/input.pl